How to Set Up a ¹H–¹⁵N HSQC Experiment in TopSpin
This tutorial walks you through every step—from loading your sample to fine-tuning shims and measuring signal-to-noise—so you can acquire high-quality HSQC data and compare the classic and BEST-HSQC variants. There is another tutorial based on a workshop, which will be eventually merged to this one.
1. Prepare a New Experiment Folder
- Copy a previously calibrated ¹H experiment
- In TopSpin, select an earlier proton pulse-calibration experiment (e.g.
zg) and pressnew. - Create a new dataset and copy the parameters into it.
- In TopSpin, select an earlier proton pulse-calibration experiment (e.g.
- Set the temperature
- Press
edteand enter the desired temperature.
- Press
-
Insert the sample into the spectrometer.
- Lock on the solvent
- Press
lock, choose the correct solvent (e.g.H2O+D2O), and wait ≈ 10 min for temperature stabilization.
- Press
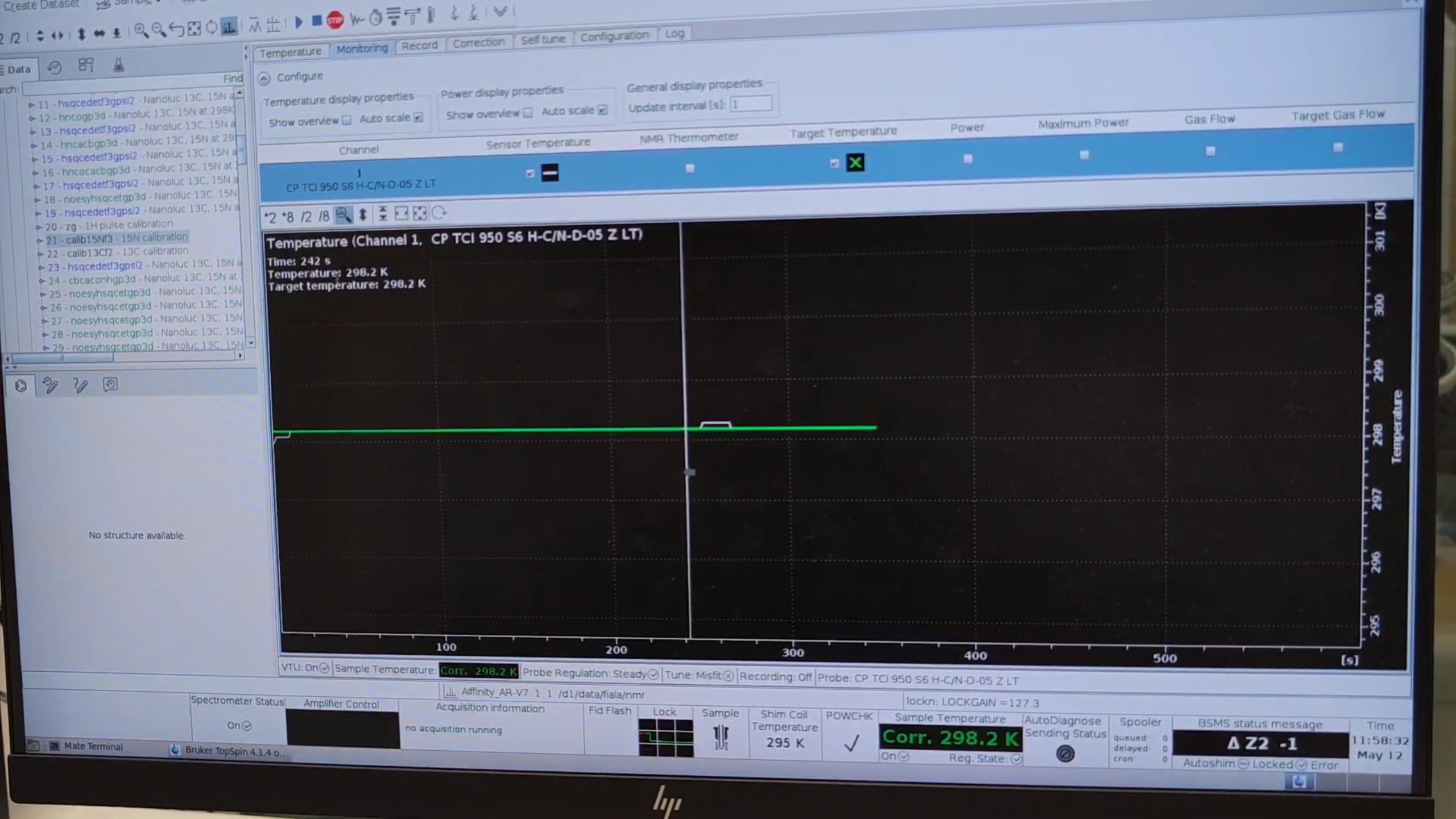
Tip: While the sample equilibrates, you can start configuring the HSQC experiment.
2. Load and Configure the HSQC Pulse Program
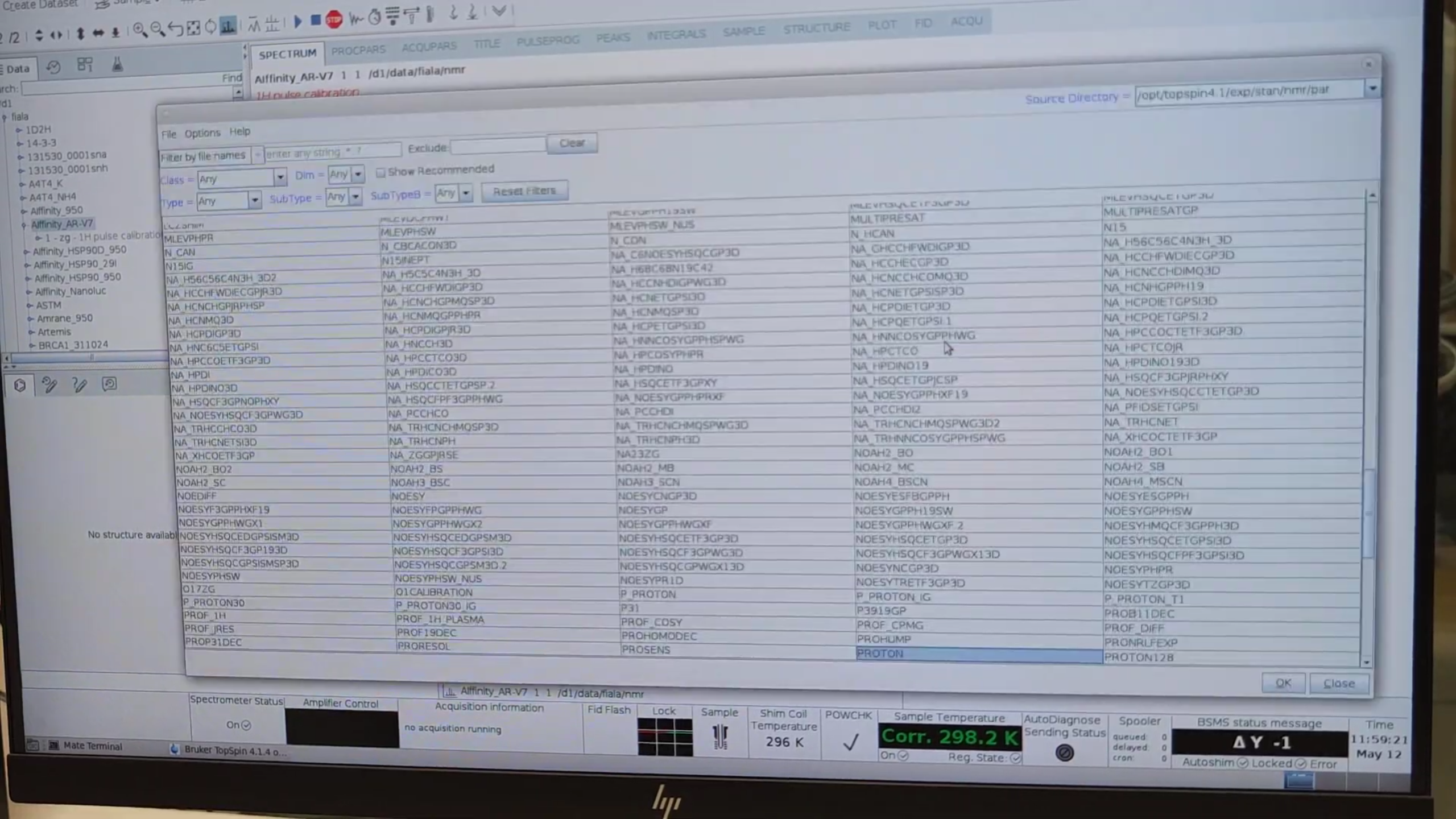
- Press
new→ Read parameter set → choose an HSQC sequence (e.g.HSQCETF3FGPSI). - Select Execute getprosol to import pulse-length/power values.
-
Enter a short description under Title and click OK.
- Verify acquisition channels
- Press
edasp.
- Press
- Set basic parameters
- Press
edaand adjust- SW (spectral width)
- O1P (reference offset)
- AQ (acquisition time)
- TD / DS / NS as needed for total experiment time (
expt).
- Press
3. Probe Tuning & Matching
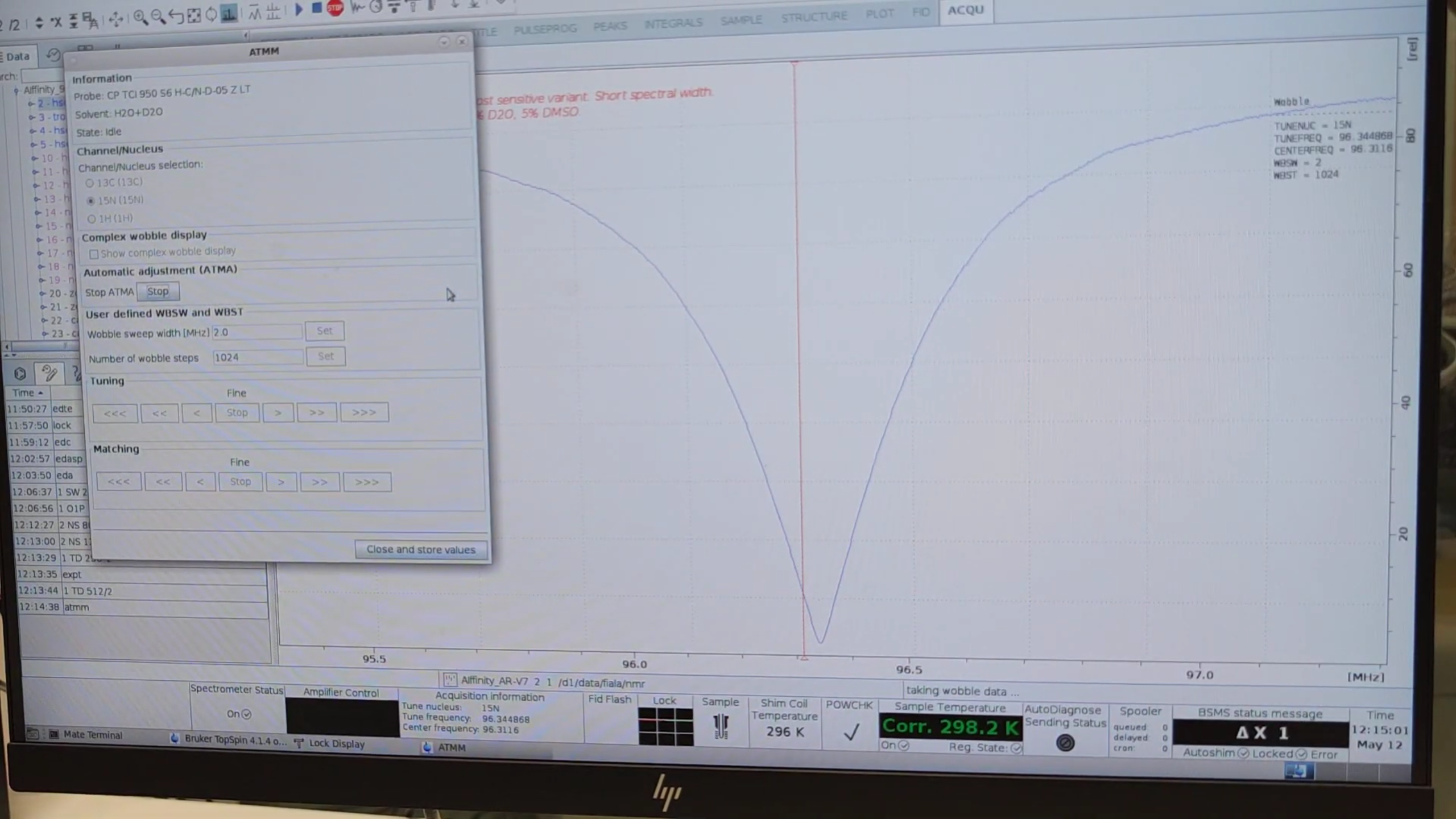
- Press
atmm(manual tune/match). - Select
15N, click Start, then repeat for1H. -
When automatic tuning finishes, switch to manual mode and center the minima on both channels.
- Press
loopadjto refine lock parameters.
4. Automated Shimming (Shigemi Tube)
- Launch the GUI:
topshim gui.
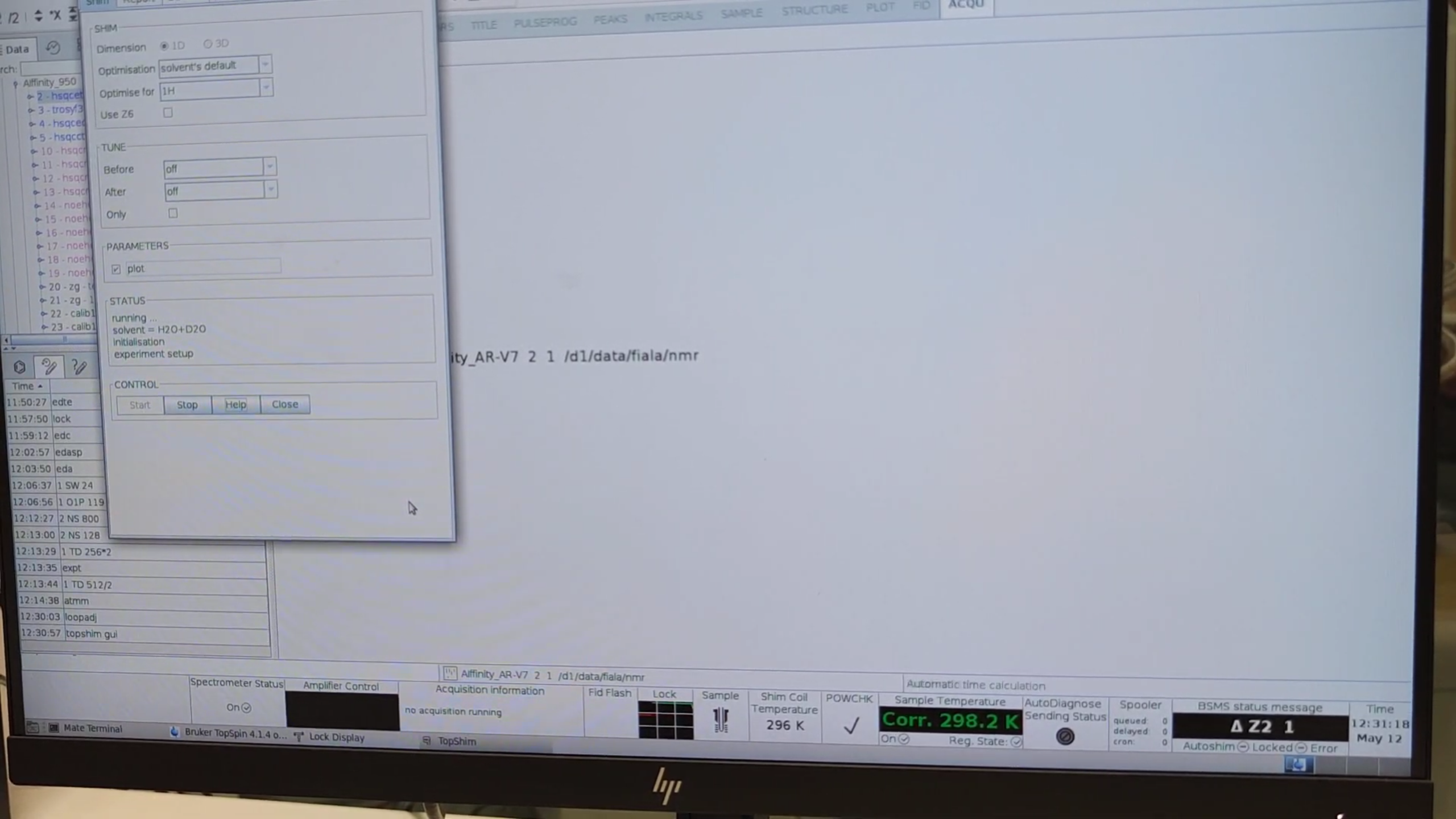
- Under PARAMETERS type
plotand start an initial shim. - Examine the field profile:
TopshimData → 1D_maps_field → zg30(see liquid range).
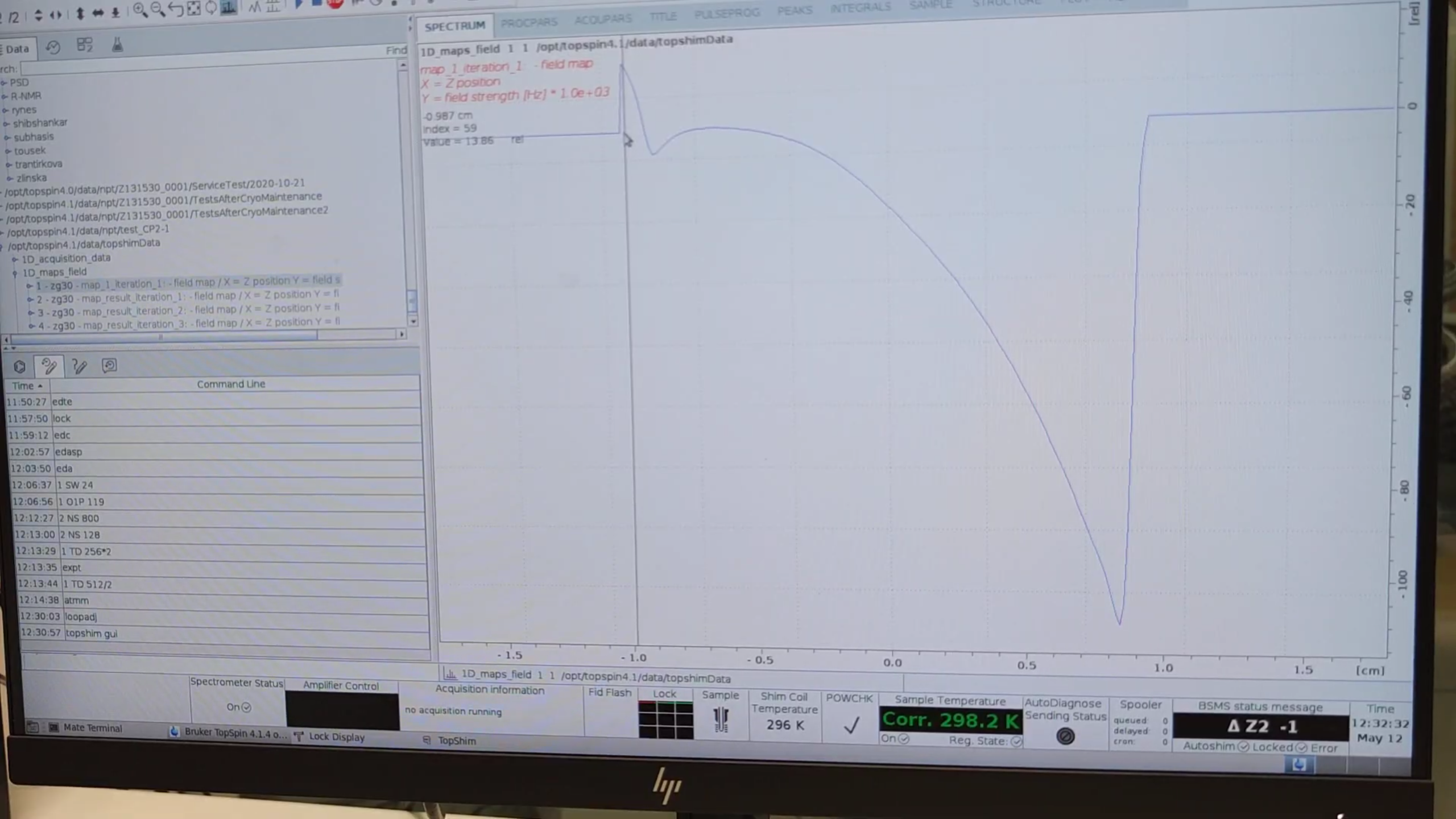
- Run a 3D shim:
topshim gui→ choose 3D, enable Use Z6.- Set Before / After to
Z-X-Y-XZ-YZ-Z. - Under PARAMETERS enter the z-range you measured, e.g.
zrange=-0.881,0.84, then click Start.
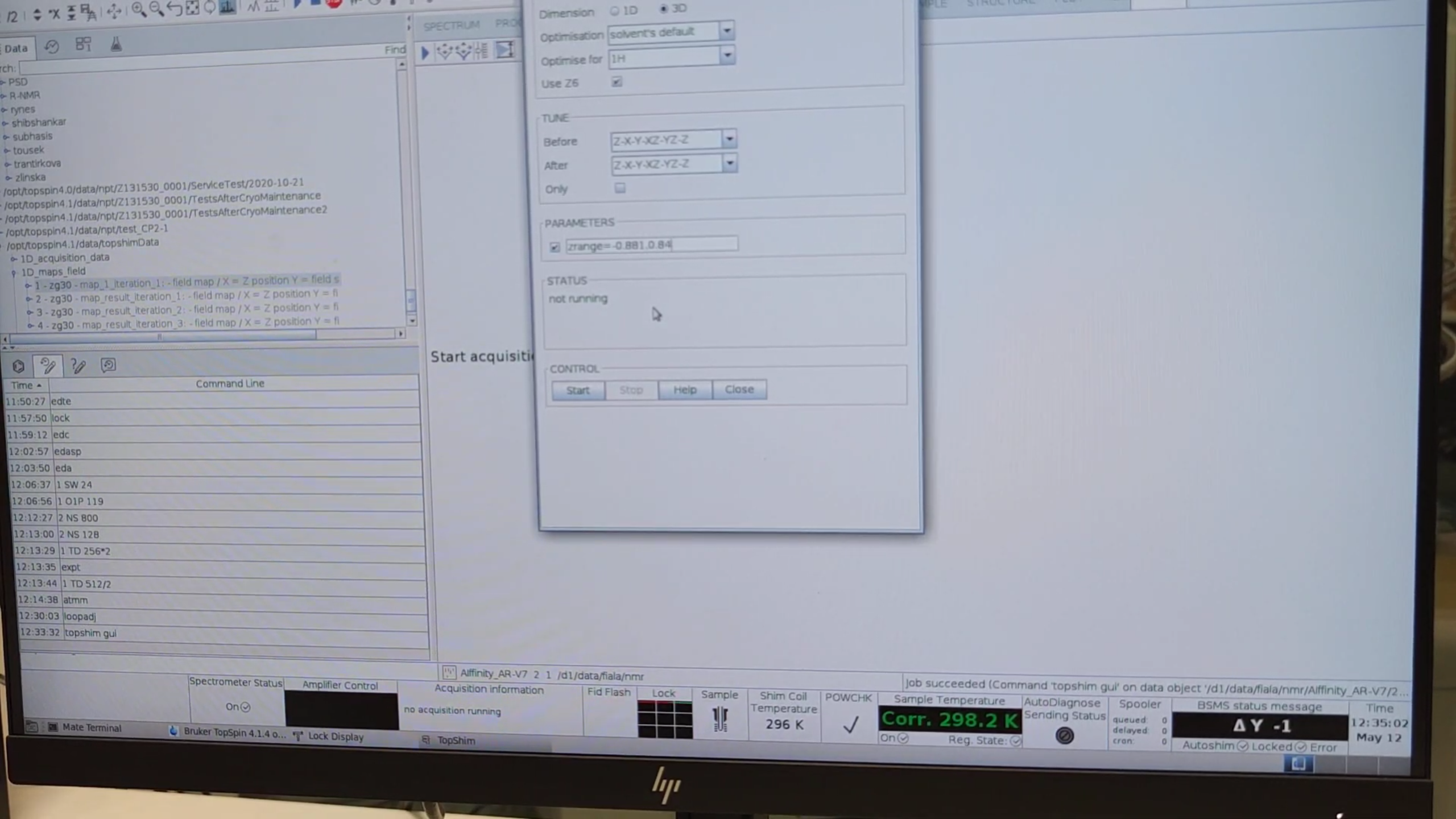
5. Fine-Tuning the ¹H Pulse Length
The following commands were issued in the console (no screenshots):
lockdisp
loopadj
wsh
bsmsdisp
re 1 # recall the 1D proton dataset
eda
1 AQ 0.524 # extend acquisition time
1 TD 64K
pulsecalc
ased
1 P1 1
zg # acquire 1D ¹H
gm
ft
mc
pp
Calibrate P1
- Open
ased, set1 P1 15.59*4. - Acquire (
zg) the 1D proton spectrum. fp→ Check phasing; if lock drifted (DMSO present), re-lock..ph→ manual phase → save.-
Repeat
zgto iterate pulse calibration:- Goal: equal positive and negative components (ideally zero net signal).
- If more negative, increase P1 via
ased(1 P1 62.520) and re-runzg. - Continue until balanced.
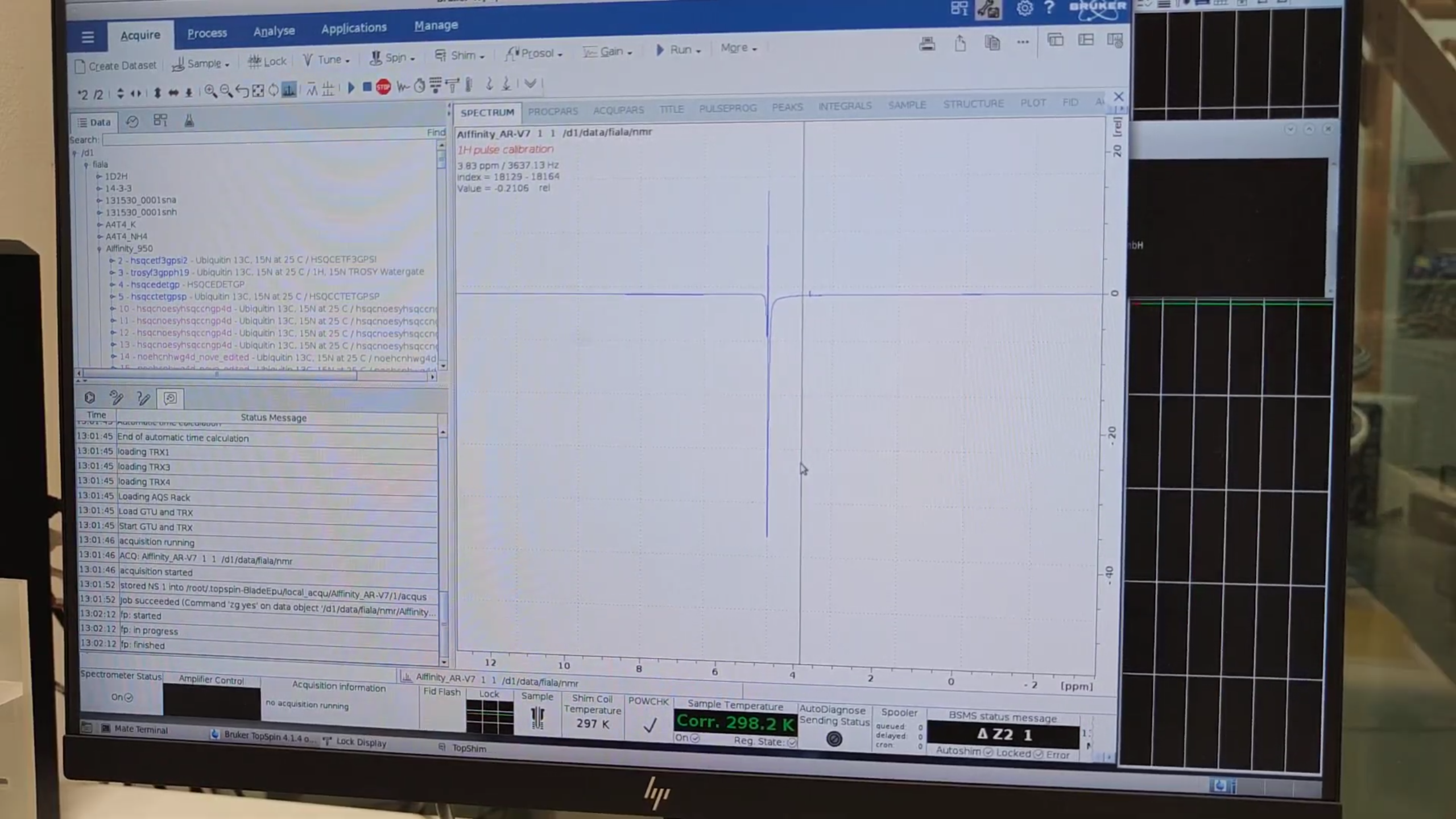
- Divide the final P1 by 4 (90° pulse) in
eda.
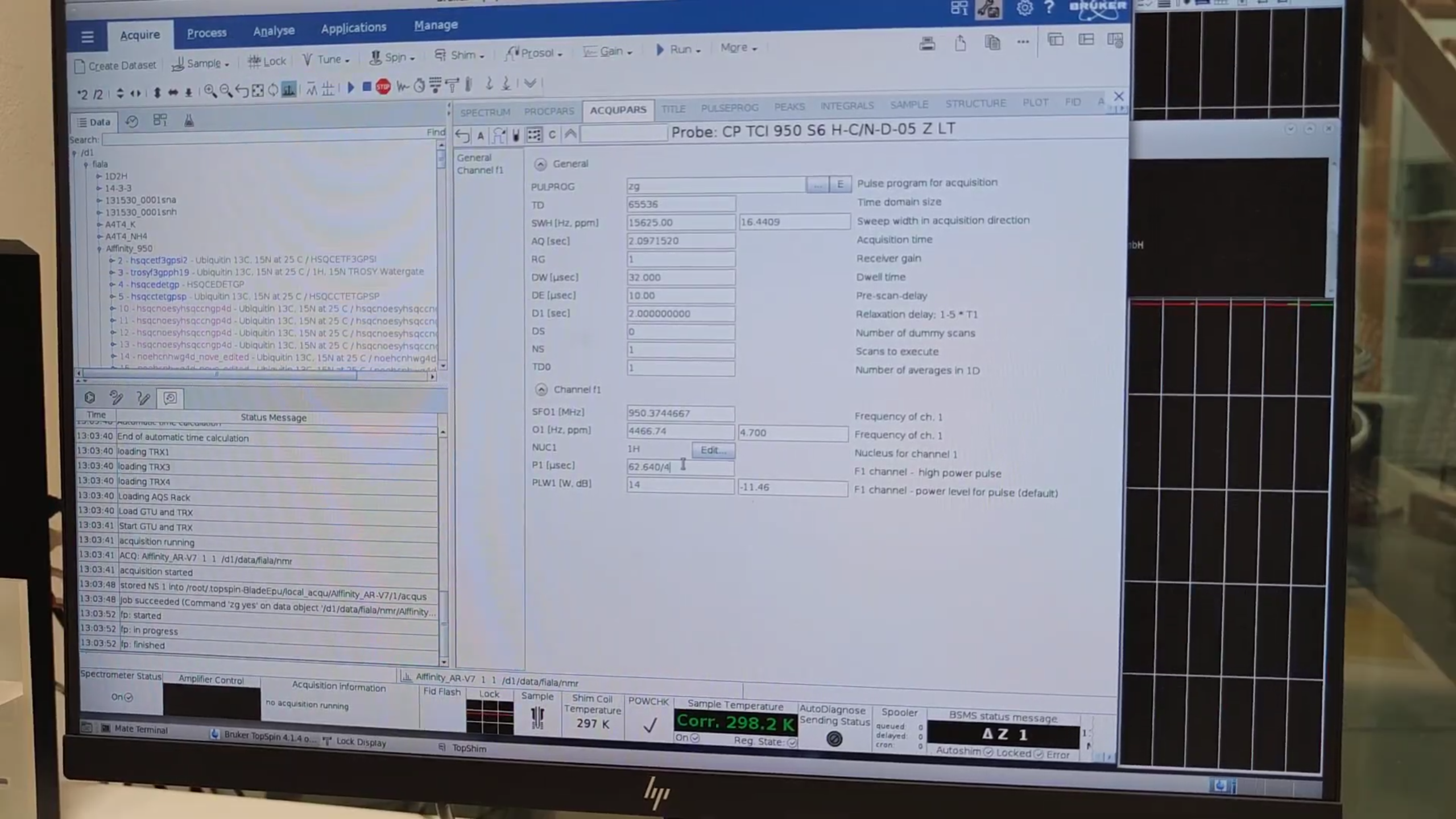
6. Water Referencing & Final HSQC Setup
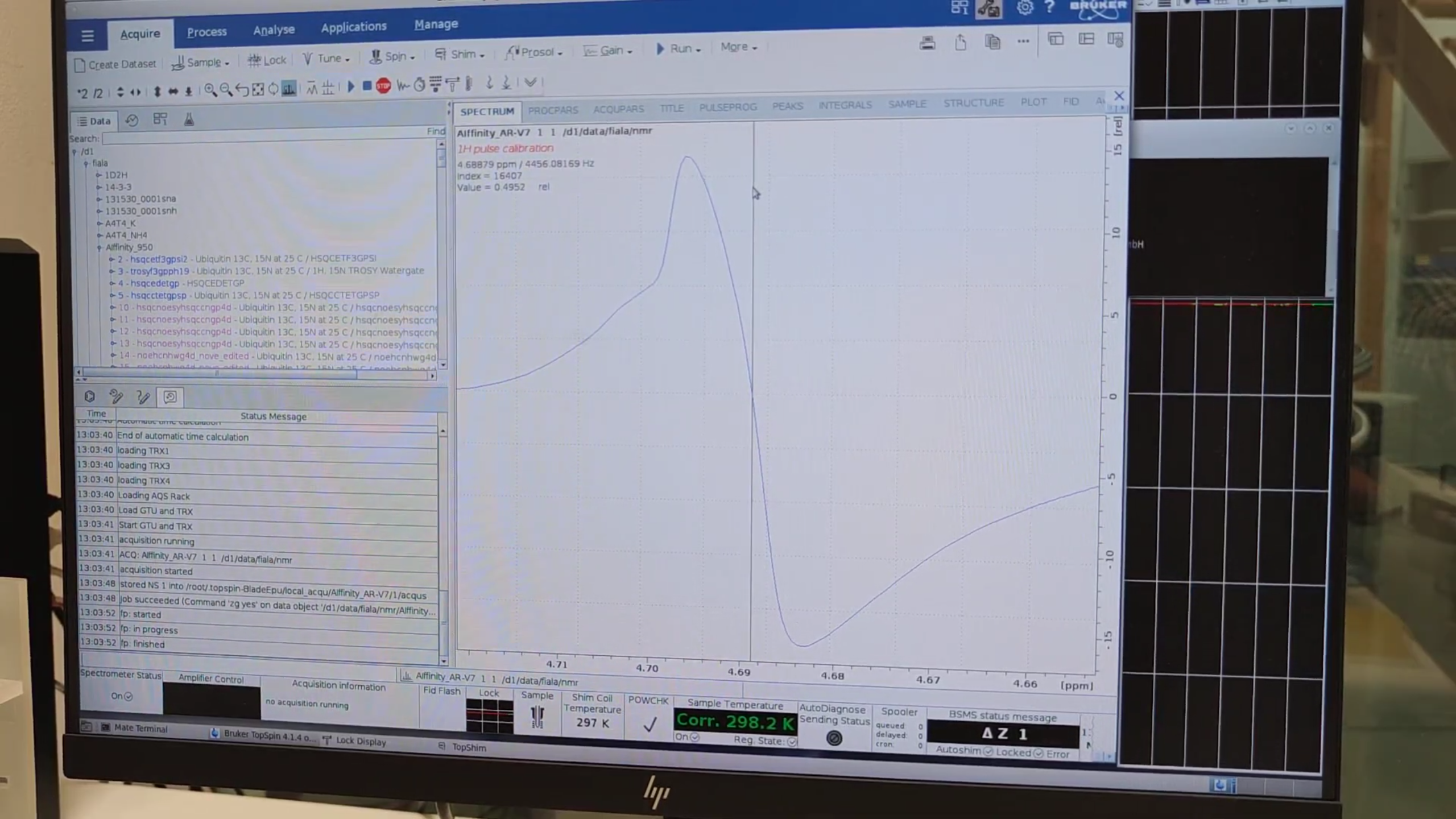
-
Zoom into the water peak, note the zero-intensity carrier (e.g. 4456.08 Hz).
-
re 2→ return to HSQC dataset →eda→ set O1{F2} to that value. -
Execute
getprosol 1H 15.66 14W(15.66 µs length of 90 degree pulse, PLW1{F2}=14 W power). -
Verify in
ased, then start the 2D HSQC:zg. -
While acquiring, extract the 1D strip:
qsin→ OK → peaks between 8–10 ppm confirm protein visibility.- Phase with
.phif needed.
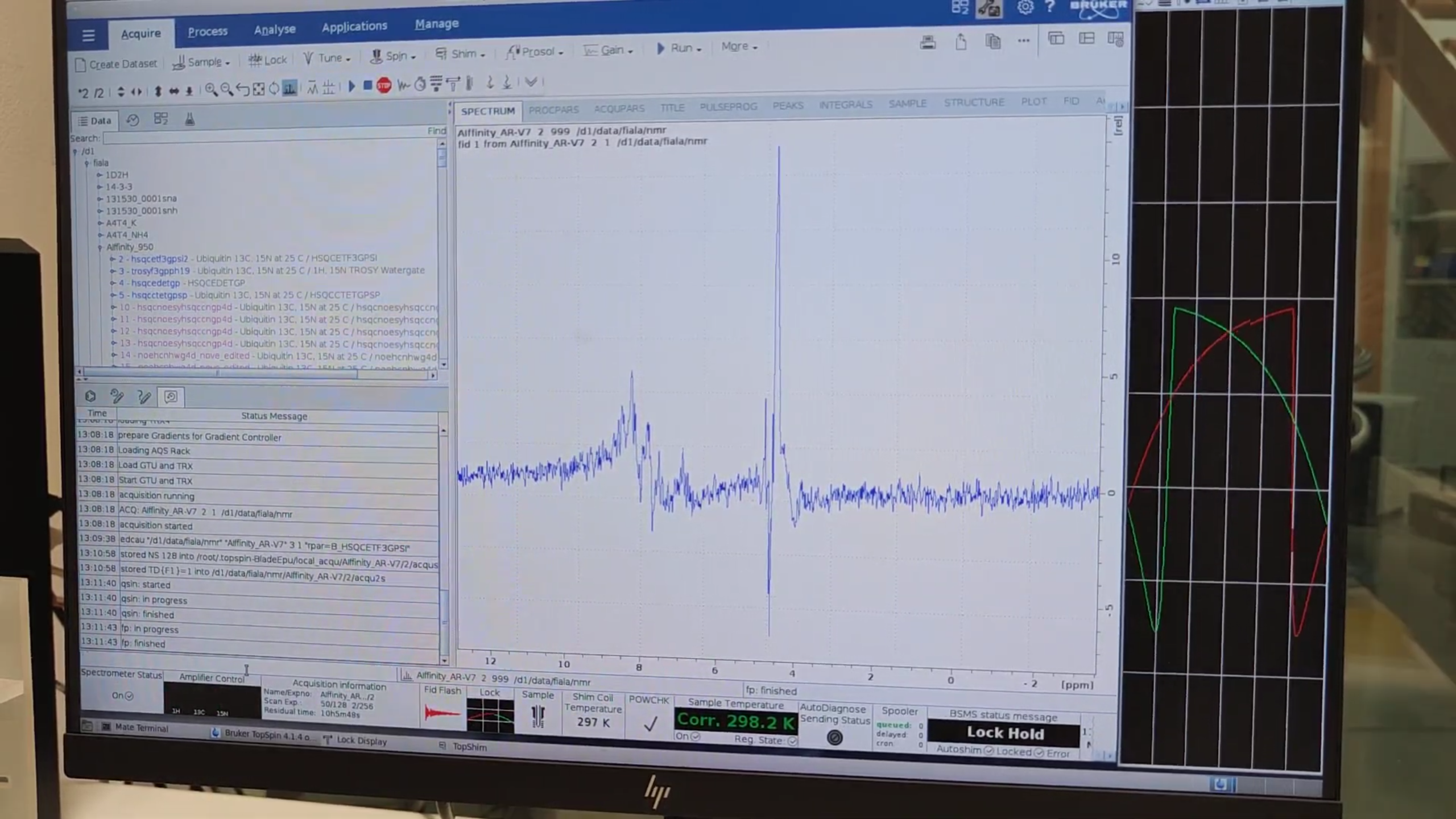
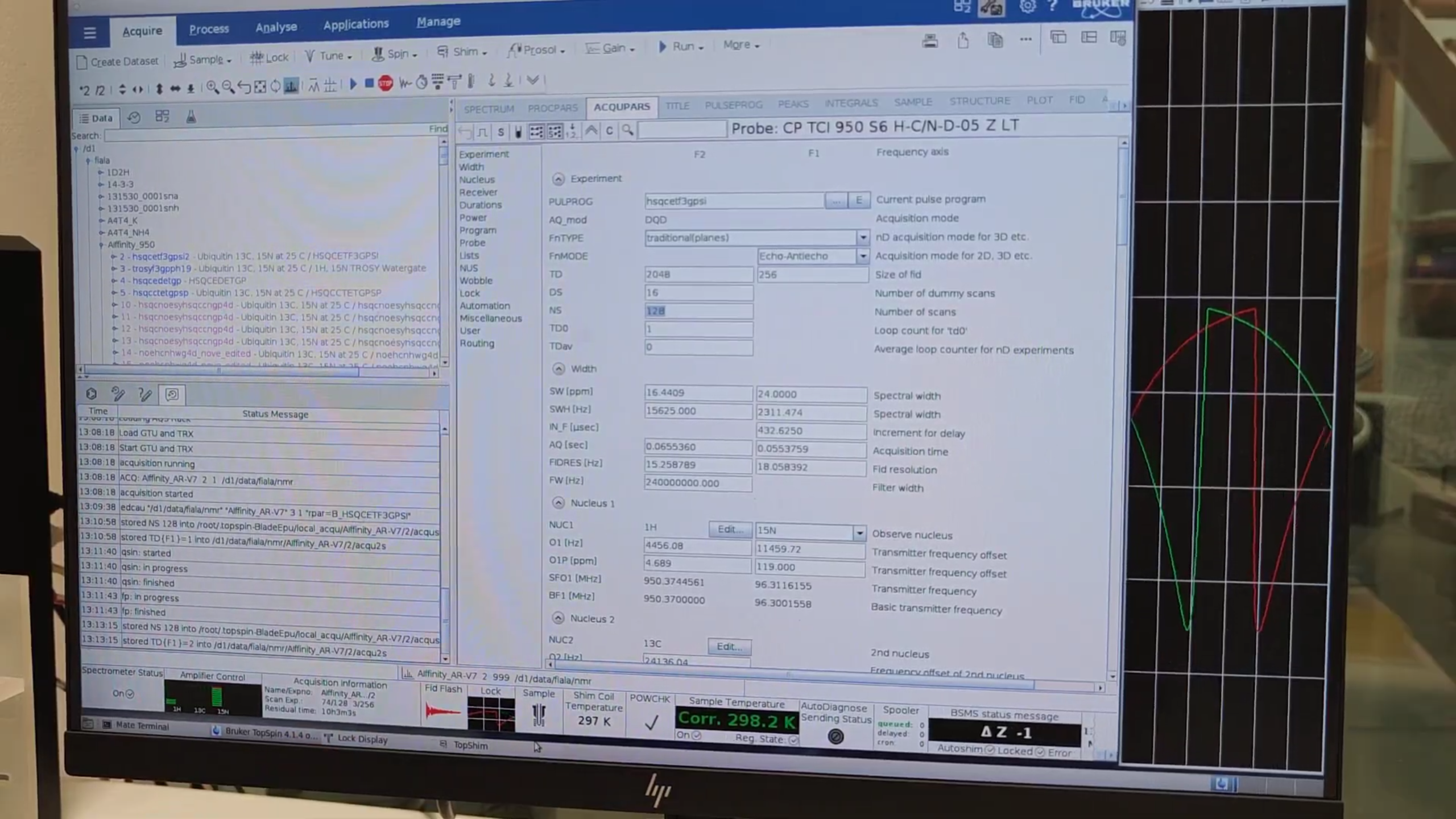
7. Setting Up the BEST-HSQC
- Press
new→ Read parameters → selectB_HSQCETF3GPSI(BEST variant). -
In
eda, note the shorter TD{F1} (faster but lower resolution).- Adjust TD, SW, NS to balance time vs. resolution.
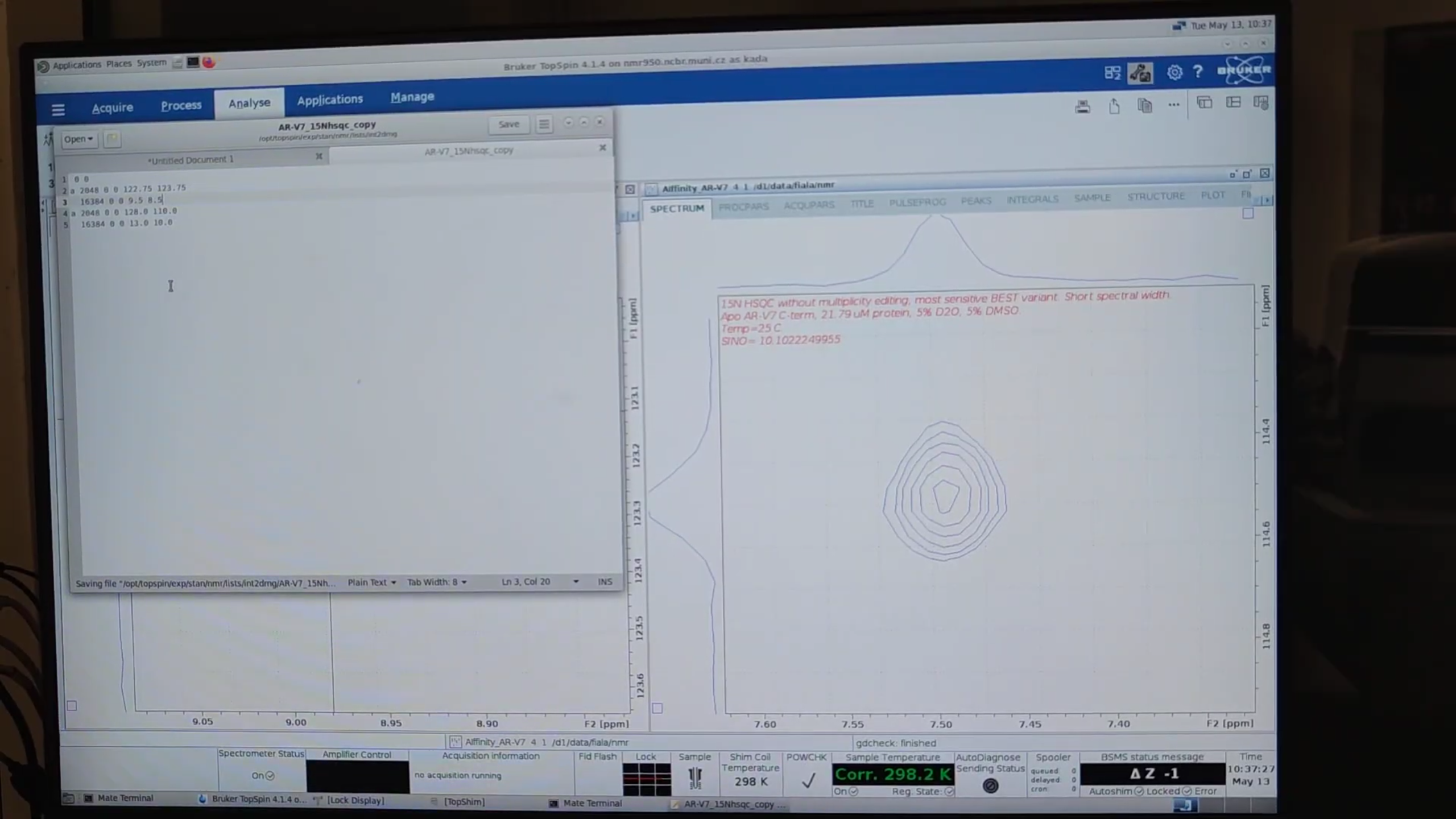
8. Measuring 2D Signal-to-Noise (Classic vs. BEST)
- Display both processed datasets side-by-side.
xfb→ Fourier transform →abs2thenabs1for baseline correction.- Synchronize views, pick a strong peak, record its region.
-
Create an
int2drngfile (peak1.txt) with:0 0 a 2048 0 0 122.75 123.75 16384 0 0 9.5 8.5 a 1024 0 0 128.0 110.0 16384 0 0 13.0 10.4
- Run
sino2D, choose peak1.txt—note SINO value. - Repeat for the other dataset and for 2–3 additional peaks to compare classic vs. BEST performance.
9. Summary Checklist
| Step | Command | Purpose |
|---|---|---|
| New dataset | new |
Copy parameters |
| Temperature | edte |
Set temp |
| Parameter check | edasp, eda |
Verify & edit settings |
| Probe tune/match | atmm |
¹H & ¹⁵N matching |
| Lock optimize | loopadj |
Fine lock |
| Shim | topshim gui |
1-D then 3-D shim |
| Pulse calibration | zg, ased |
Optimize P1 |
| Water reference | eda |
Set O1{F2} |
| Acquire HSQC | zg |
Start 2D run |
| BEST HSQC | new → BEST sequence |
Faster variant |
| S/N analysis | sino2D |
Compare datasets |
Authors
- Thomas Evangelidis